
Coreform Cubit provided an intuitive platform for meshing the model geometry, especially in complex biological tissues with fine anatomical detail. Without Cubit’s capabilities, many important features would be lost in coarser, simplified anatomical models.
Nicholas Glover, Ph.D. Student
University at Buffalo
Background
Nicholas Glover is a Ph.D. student in the Systems Biomedicine and Pharmaceutics Laboratory led by Dr. Ashlee N. Ford Versypt, where he works on the mechanisms of chemical and biomechanical damage in kidney tissues during diabetic kidney disease. Nicholas studied the biomechanical properties of the glomerulus, the kidney’s major anatomical structure responsible for filtering waste from the blood, which is the primary target of diabetic tissue damage. He sought to establish a connection between morphological changes in the anatomical structure resulting from disease and changes in the transport of filtrate as it moves through the glomerular filtration barrier, to better understand the progression of diabetic kidney damage. To simulate these processes in a computational model, ANSYS fluent and FEbio have been used as dedicated solvers.
The project integrates several processes, each shown individually to contribute to kidney injury in diabetes, into a sophisticated computer simulation. This computational tool will aid in understanding how the processes interact and how diabetic kidney damage begins and changes over time. In the long term, results from this project will help predict the impacts of many competing factors on kidney health during diabetes management. The results of these simulations can give an estimation of the filtering capacity of each layer in a quantitative way. This information can be used to guide effective treatment plans for patients and to predict and monitor disease states in a non-invasive manner.
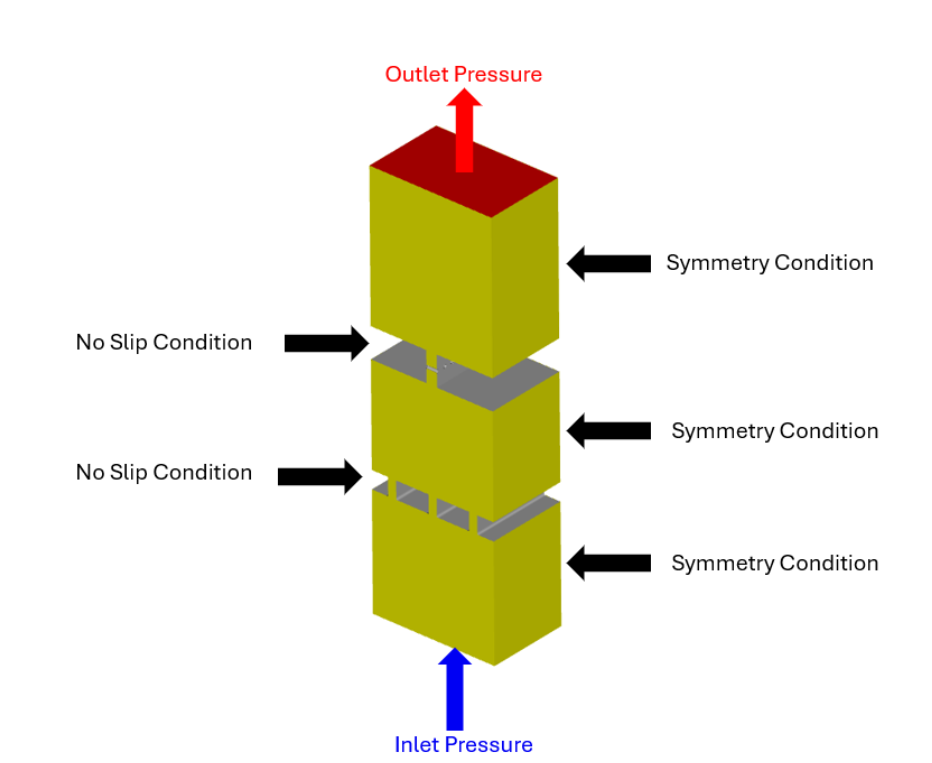
Figure 1: Geometry of the kidney’s glomerular filtration barrier and the boundary conditions associated with each surface to be meshed in Coreform Cubit. The vertically stacked layers of the tissue represented here each have different porous media material properties.
THE PROBLEM
As the kidney’s function to filter waste from the blood depends on the physical filtering layers, a 3D representative geometry of the filtering structure is first needed to simulate the filtration process. To build such a geometry of the system, biological images of kidney glomeruli were used to construct multiple disease-state geometries using structure parameters derived from the images. To obtain an accurate CFD model of the layered filtration barrier, meshing software capable of producing high-quality meshes was required. Some layers are the gaps between biological cells of different sizes and shapes, while others are more amorphous, porous materials. So it is important to also be able to create element sets that correspond to the various biological materials present in the model across the different layers.
The researchers introduced small variations in the filtration barrier’s fine-scale anatomy to study how specific structural features influence local fluid flow. Because these anatomical variants were created with parametric CAD models, the workflow required automated meshing to support efficient and repeatable analysis across changing geometries (Figure 1).
Other available software lacked the meshing tools needed to accommodate the complex glomerulus structure and often had user interfaces that made the workflow less intuitive.
THE SOLUTION
Previously published anatomical measurements were used to define the geometry for the solute transport study through the glomerular filtration barrier. From these measurements, the team constructed a layered representation of the glomerular filtration barrier, capturing the fluid channels and porous regions through which filtrate passes. Under normal conditions, this filtrate includes water and small molecules; in disease states, it can also include proteins. To support the ANSYS Fluent multiphysics simulations, this geometry required a mesh that could accurately resolve the thin-layered structure, preserve fine geometric detail, and maintain the boundary information needed to represent contact between the filtrate and the cell surfaces that form the filtration channels. Coreform Cubit was used to generate this mesh (see Figure 2), providing the level of geometric fidelity and boundary definition required for the analysis.
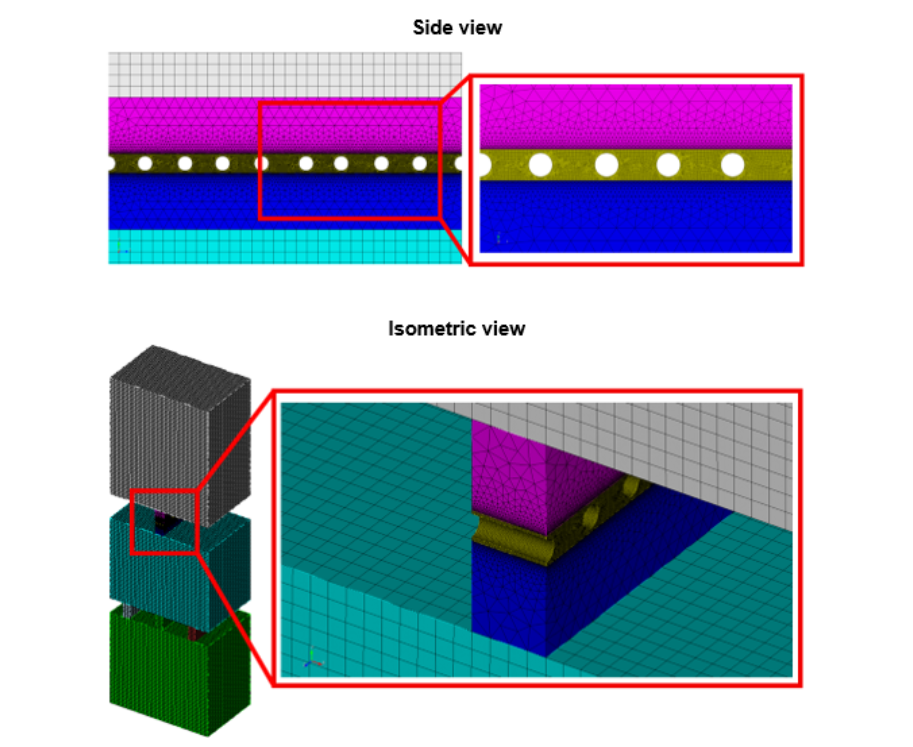
Figure 2: High-quality mesh for the kidney’s glomerular filtration barrier produced by Coreform Cubit. The meshing captured fine features of the narrow channels, allowed biasing near the domain boundaries, and had coarser resolution in the middle of the large fluid domains.
Using previously published theoretical formulations for multiphasic transport in the kidney, the researchers assigned physical properties to the different layers and selected interface conditions that best represented physiological kidney behavior. The resulting model included fluid and biphasic domains, a normal fluid velocity based on a single-nephron glomerular filtration rate, and boundary conditions governing fluid behavior at cell-filtrate interfaces. As the project advances and additional biological variations are incorporated, Coreform Cubit’s Sculpt technology offers an efficient path for generating high-quality hexahedral meshes for both normal and diseased geometries.
In ANSYS, the team used journal-file-based workflows to run CFD simulations on the Coreform Cubit mesh (see Figures 3 and 4). These simulations were used to study hydraulic conductivity and wall shear stress in normal and diseased states, yielding distributions of fluid velocity and shearing forces along the surface of the slit diaphragm, a key filtering layer of the kidney.
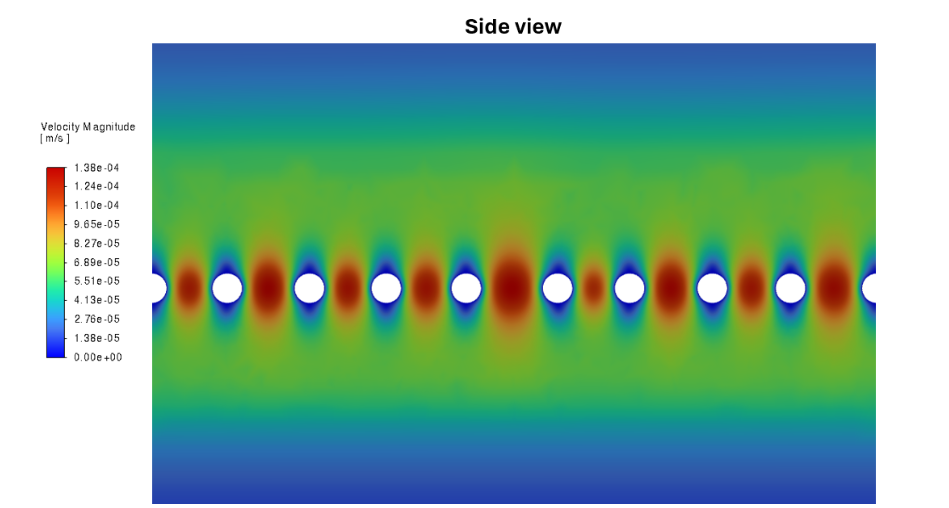
Figure 3: Fluid velocity along the surface of the slit diaphragm (the inset region of Figure 2).
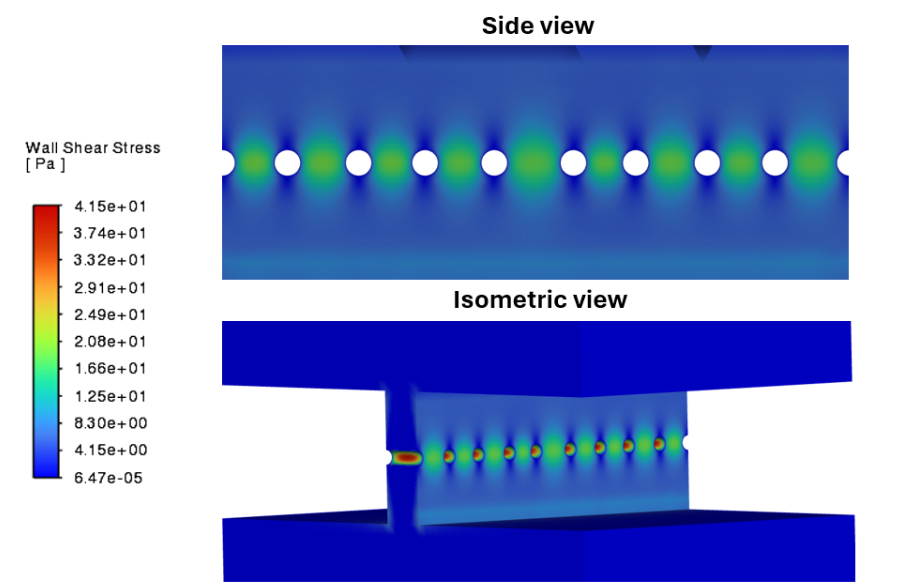
Figure 4: Wall shear stress along the surface of the slit diaphragm (the inset region of Figure 2).
CONCLUSION
Dr. Ford Versypt’s lab is developing a multiscale virtual-kidney model to study the onset and progression of diabetic kidney disease. Before Nicholas Glover’s contribution, the project focused primarily on chemical and biological processes. His work added the physical filtration barriers, creating a foundation for connecting changes in kidney structure and material behavior to the biomechanics that govern filtration.
Coreform Cubit played a central role in enabling this work. By providing the high-quality meshes needed to represent the kidney’s fine anatomical layers and integrate them into the simulation workflow, Coreform Cubit allowed the team to study how wall shear stress contributes to cell damage in a primary filtering layer and how structural changes affect protein transport across the filtration barrier. These are key mechanisms in disease progression and are directly relevant to the protein leakage used to monitor kidney damage clinically. With Coreform Cubit supporting this workflow, the team is building more predictive computational tools for understanding disease progression before permanent damage is clinically detected.














